algorithms.statistics.models.model¶
Module: algorithms.statistics.models.model¶
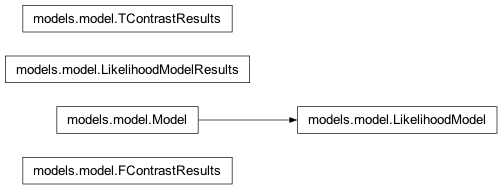
Inheritance diagram for nipy.algorithms.statistics.models.model:
Classes¶
FContrastResults¶
- class nipy.algorithms.statistics.models.model.FContrastResults(effect, covariance, F, df_num, df_den=None)¶
Bases:
objectResults from an F contrast of coefficients in a parametric model.
The class does nothing, it is a container for the results from F contrasts, and returns the F-statistics when np.asarray is called.
- __init__(effect, covariance, F, df_num, df_den=None)¶
LikelihoodModel¶
- class nipy.algorithms.statistics.models.model.LikelihoodModel¶
Bases:
Model- __init__()¶
- fit()¶
Fit a model to data.
- information(theta, nuisance=None)¶
Fisher information matrix
The inverse of the expected value of
- d^2 logL / dtheta^2.
- initialize()¶
Initialize (possibly re-initialize) a Model instance.
For instance, the design matrix of a linear model may change and some things must be recomputed.
- logL(theta, Y, nuisance=None)¶
Log-likelihood of model.
- predict(design=None)¶
After a model has been fit, results are (assumed to be) stored in self.results, which itself should have a predict method.
- score(theta, Y, nuisance=None)¶
Gradient of logL with respect to theta.
This is the score function of the model
LikelihoodModelResults¶
- class nipy.algorithms.statistics.models.model.LikelihoodModelResults(theta, Y, model, cov=None, dispersion=1.0, nuisance=None, rank=None)¶
Bases:
objectClass to contain results from likelihood models
- __init__(theta, Y, model, cov=None, dispersion=1.0, nuisance=None, rank=None)¶
Set up results structure
- Parameters:
- thetandarray
parameter estimates from estimated model
- Yndarray
data
- model
LikelihoodModelinstance model used to generate fit
- covNone or ndarray, optional
covariance of thetas
- dispersionscalar, optional
multiplicative factor in front of cov
- nuisanceNone of ndarray
parameter estimates needed to compute logL
- rankNone or scalar
rank of the model. If rank is not None, it is used for df_model instead of the usual counting of parameters.
Notes
The covariance of thetas is given by:
dispersion * cov
For (some subset of models) dispersion will typically be the mean square error from the estimated model (sigma^2)
- AIC()¶
Akaike Information Criterion
- BIC()¶
Schwarz’s Bayesian Information Criterion
- Fcontrast(matrix, dispersion=None, invcov=None)¶
Compute an Fcontrast for a contrast matrix matrix.
Here, matrix M is assumed to be non-singular. More precisely
\[M pX pX' M'\]is assumed invertible. Here, \(pX\) is the generalized inverse of the design matrix of the model. There can be problems in non-OLS models where the rank of the covariance of the noise is not full.
See the contrast module to see how to specify contrasts. In particular, the matrices from these contrasts will always be non-singular in the sense above.
- Parameters:
- matrix1D array-like
contrast matrix
- dispersionNone or float, optional
If None, use
self.dispersion- invcovNone or array, optional
Known inverse of variance covariance matrix. If None, calculate this matrix.
- Returns:
- f_res
FContrastResultsinstance with attributes F, df_den, df_num
- f_res
Notes
For F contrasts, we now specify an effect and covariance
- Tcontrast(matrix, store=('t', 'effect', 'sd'), dispersion=None)¶
Compute a Tcontrast for a row vector matrix
To get the t-statistic for a single column, use the ‘t’ method.
- Parameters:
- matrix1D array-like
contrast matrix
- storesequence, optional
components of t to store in results output object. Defaults to all components (‘t’, ‘effect’, ‘sd’).
- dispersionNone or float, optional
- Returns:
- res
TContrastResultsobject
- res
- conf_int(alpha=0.05, cols=None, dispersion=None)¶
The confidence interval of the specified theta estimates.
- Parameters:
- alphafloat, optional
The alpha level for the confidence interval. ie., alpha = .05 returns a 95% confidence interval.
- colstuple, optional
cols specifies which confidence intervals to return
- dispersionNone or scalar
scale factor for the variance / covariance (see class docstring and
vcovmethod docstring)
- Returns:
- cisndarray
cis is shape
(len(cols), 2)where each row contains [lower, upper] for the given entry in cols
Notes
Confidence intervals are two-tailed. TODO: tails : string, optional
tails can be “two”, “upper”, or “lower”
Examples
>>> from numpy.random import standard_normal as stan >>> from nipy.algorithms.statistics.models.regression import OLSModel >>> x = np.hstack((stan((30,1)),stan((30,1)),stan((30,1)))) >>> beta=np.array([3.25, 1.5, 7.0]) >>> y = np.dot(x,beta) + stan((30)) >>> model = OLSModel(x).fit(y) >>> confidence_intervals = model.conf_int(cols=(1,2))
- logL()¶
The maximized log-likelihood
- t(column=None)¶
Return the (Wald) t-statistic for a given parameter estimate.
Use Tcontrast for more complicated (Wald) t-statistics.
- vcov(matrix=None, column=None, dispersion=None, other=None)¶
Variance/covariance matrix of linear contrast
- Parameters:
- matrix: (dim, self.theta.shape[0]) array, optional
numerical contrast specification, where
dimrefers to the ‘dimension’ of the contrast i.e. 1 for t contrasts, 1 or more for F contrasts.- column: int, optional
alternative way of specifying contrasts (column index)
- dispersion: float or (n_voxels,) array, optional
value(s) for the dispersion parameters
- other: (dim, self.theta.shape[0]) array, optional
alternative contrast specification (?)
- Returns:
- cov: (dim, dim) or (n_voxels, dim, dim) array
the estimated covariance matrix/matrices
- Returns the variance/covariance matrix of a linear contrast of the
- estimates of theta, multiplied by dispersion which will often be an
- estimate of dispersion, like, sigma^2.
- The covariance of interest is either specified as a (set of) column(s)
- or a matrix.
Model¶
- class nipy.algorithms.statistics.models.model.Model¶
Bases:
objectA (predictive) statistical model.
The class Model itself does nothing but lays out the methods expected of any subclass.
- __init__()¶
- fit()¶
Fit a model to data.
- initialize()¶
Initialize (possibly re-initialize) a Model instance.
For instance, the design matrix of a linear model may change and some things must be recomputed.
- predict(design=None)¶
After a model has been fit, results are (assumed to be) stored in self.results, which itself should have a predict method.
TContrastResults¶
- class nipy.algorithms.statistics.models.model.TContrastResults(t, sd, effect, df_den=None)¶
Bases:
objectResults from a t contrast of coefficients in a parametric model.
The class does nothing, it is a container for the results from T contrasts, and returns the T-statistics when np.asarray is called.
- __init__(t, sd, effect, df_den=None)¶
