algorithms.statistics.models.regression¶
Module: algorithms.statistics.models.regression¶
Inheritance diagram for nipy.algorithms.statistics.models.regression:
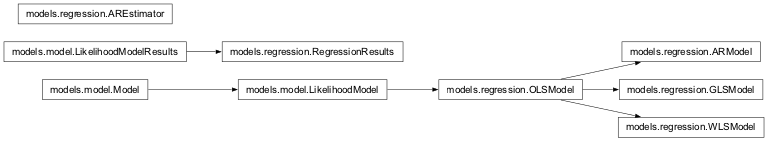
This module implements some standard regression models: OLS and WLS models, as well as an AR(p) regression model.
Models are specified with a design matrix and are fit using their ‘fit’ method.
Subclasses that have more complicated covariance matrices should write over the ‘whiten’ method as the fit method prewhitens the response by calling ‘whiten’.
General reference for regression models:
- ‘Introduction to Linear Regression Analysis’, Douglas C. Montgomery,
Elizabeth A. Peck, G. Geoffrey Vining. Wiley, 2006.
Classes¶
AREstimator¶
- class nipy.algorithms.statistics.models.regression.AREstimator(model, p=1)¶
Bases:
objectA class to estimate AR(p) coefficients from residuals
- __init__(model, p=1)¶
Bias-correcting AR estimation class
- Parameters:
- model
OSLModelinstance A models.regression.OLSmodel instance, where model has attribute
design- pint, optional
Order of AR(p) noise
- model
ARModel¶
- class nipy.algorithms.statistics.models.regression.ARModel(design, rho)¶
Bases:
OLSModelA regression model with an AR(p) covariance structure.
In terms of a LikelihoodModel, the parameters are beta, the usual regression parameters, and sigma, a scalar nuisance parameter that shows up as multiplier in front of the AR(p) covariance.
- The linear autoregressive process of order p–AR(p)–is defined as:
TODO
Examples
>>> from nipy.algorithms.statistics.api import Term, Formula >>> data = np.rec.fromarrays(([1,3,4,5,8,10,9], range(1,8)), ... names=('Y', 'X')) >>> f = Formula([Term("X"), 1]) >>> dmtx = f.design(data, return_float=True) >>> model = ARModel(dmtx, 2)
We go through the
model.iterative_fitprocedure long-hand:>>> for i in range(6): ... results = model.fit(data['Y']) ... print("AR coefficients:", model.rho) ... rho, sigma = yule_walker(data["Y"] - results.predicted, ... order=2, ... df=model.df_resid) ... model = ARModel(model.design, rho) ... AR coefficients: [ 0. 0.] AR coefficients: [-0.61530877 -1.01542645] AR coefficients: [-0.72660832 -1.06201457] AR coefficients: [-0.7220361 -1.05365352] AR coefficients: [-0.72229201 -1.05408193] AR coefficients: [-0.722278 -1.05405838] >>> results.theta array([ 1.59564228, -0.58562172]) >>> results.t() array([ 38.0890515 , -3.45429252]) >>> print(results.Tcontrast([0,1])) <T contrast: effect=-0.58562172384377043, sd=0.16953449108110835, t=-3.4542925165805847, df_den=5> >>> print(results.Fcontrast(np.identity(2))) <F contrast: F=4216.810299725842, df_den=5, df_num=2>
Reinitialize the model, and do the automated iterative fit
>>> model.rho = np.array([0,0]) >>> model.iterative_fit(data['Y'], niter=3) >>> print(model.rho) [-0.7220361 -1.05365352]
- __init__(design, rho)¶
Initialize AR model instance
- Parameters:
- designndarray
2D array with design matrix
- rhoint or array-like
If int, gives order of model, and initializes rho to zeros. If ndarray, gives initial estimate of rho. Be careful as
ARModel(X, 1) != ARModel(X, 1.0).
- fit(Y)¶
Fit model to data Y
Full fit of the model including estimate of covariance matrix, (whitened) residuals and scale.
- Parameters:
- Yarray-like
The dependent variable for the Least Squares problem.
- Returns:
- fitRegressionResults
- has_intercept()¶
Check if column of 1s is in column space of design
- information(beta, nuisance=None)¶
Returns the information matrix at (beta, Y, nuisance).
See logL for details.
- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- nuisancedict
A dict with key ‘sigma’, which is an estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / nwhere n=Y.shape[0], X=self.design.
- Returns:
- infoarray
The information matrix, the negative of the inverse of the Hessian of the of the log-likelihood function evaluated at (theta, Y, nuisance).
- initialize(design)¶
Initialize (possibly re-initialize) a Model instance.
For instance, the design matrix of a linear model may change and some things must be recomputed.
- iterative_fit(Y, niter=3)¶
Perform an iterative two-stage procedure to estimate AR(p) parameters and regression coefficients simultaneously.
- Parameters:
- Yndarray
data to which to fit model
- niteroptional, int
the number of iterations (default 3)
- Returns:
- None
- logL(beta, Y, nuisance=None)¶
Returns the value of the loglikelihood function at beta.
Given the whitened design matrix, the loglikelihood is evaluated at the parameter vector, beta, for the dependent variable, Y and the nuisance parameter, sigma.
- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- Yndarray
The dependent variable
- nuisancedict, optional
A dict with key ‘sigma’, which is an optional estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / n, where n=Y.shape[0], X=self.design.
- Returns:
- loglffloat
The value of the loglikelihood function.
Notes
The log-Likelihood Function is defined as
\[\ell(\beta,\sigma,Y)= -\frac{n}{2}\log(2\pi\sigma^2) - \|Y-X\beta\|^2/(2\sigma^2)\]The parameter \(\sigma\) above is what is sometimes referred to as a nuisance parameter. That is, the likelihood is considered as a function of \(\beta\), but to evaluate it, a value of \(\sigma\) is needed.
If \(\sigma\) is not provided, then its maximum likelihood estimate:
\[\hat{\sigma}(\beta) = \frac{\text{SSE}(\beta)}{n}\]is plugged in. This likelihood is now a function of only \(\beta\) and is technically referred to as a profile-likelihood.
References
[1]Green. “Econometric Analysis,” 5th ed., Pearson, 2003.
- predict(design=None)¶
After a model has been fit, results are (assumed to be) stored in self.results, which itself should have a predict method.
- rank()¶
Compute rank of design matrix
- score(beta, Y, nuisance=None)¶
Gradient of the loglikelihood function at (beta, Y, nuisance).
The graient of the loglikelihood function at (beta, Y, nuisance) is the score function.
See
logL()for details.- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- Yndarray
The dependent variable.
- nuisancedict, optional
A dict with key ‘sigma’, which is an optional estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / n, where n=Y.shape[0], X=self.design.
- Returns:
- The gradient of the loglikelihood function.
- whiten(X)¶
Whiten a series of columns according to AR(p) covariance structure
- Parameters:
- Xarray-like of shape (n_features)
array to whiten
- Returns:
- wXndarray
X whitened with order self.order AR
GLSModel¶
- class nipy.algorithms.statistics.models.regression.GLSModel(design, sigma)¶
Bases:
OLSModelGeneralized least squares model with a general covariance structure
- __init__(design, sigma)¶
- Parameters:
- designarray-like
This is your design matrix. Data are assumed to be column ordered with observations in rows.
- fit(Y)¶
Fit model to data Y
Full fit of the model including estimate of covariance matrix, (whitened) residuals and scale.
- Parameters:
- Yarray-like
The dependent variable for the Least Squares problem.
- Returns:
- fitRegressionResults
- has_intercept()¶
Check if column of 1s is in column space of design
- information(beta, nuisance=None)¶
Returns the information matrix at (beta, Y, nuisance).
See logL for details.
- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- nuisancedict
A dict with key ‘sigma’, which is an estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / nwhere n=Y.shape[0], X=self.design.
- Returns:
- infoarray
The information matrix, the negative of the inverse of the Hessian of the of the log-likelihood function evaluated at (theta, Y, nuisance).
- initialize(design)¶
Initialize (possibly re-initialize) a Model instance.
For instance, the design matrix of a linear model may change and some things must be recomputed.
- logL(beta, Y, nuisance=None)¶
Returns the value of the loglikelihood function at beta.
Given the whitened design matrix, the loglikelihood is evaluated at the parameter vector, beta, for the dependent variable, Y and the nuisance parameter, sigma.
- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- Yndarray
The dependent variable
- nuisancedict, optional
A dict with key ‘sigma’, which is an optional estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / n, where n=Y.shape[0], X=self.design.
- Returns:
- loglffloat
The value of the loglikelihood function.
Notes
The log-Likelihood Function is defined as
\[\ell(\beta,\sigma,Y)= -\frac{n}{2}\log(2\pi\sigma^2) - \|Y-X\beta\|^2/(2\sigma^2)\]The parameter \(\sigma\) above is what is sometimes referred to as a nuisance parameter. That is, the likelihood is considered as a function of \(\beta\), but to evaluate it, a value of \(\sigma\) is needed.
If \(\sigma\) is not provided, then its maximum likelihood estimate:
\[\hat{\sigma}(\beta) = \frac{\text{SSE}(\beta)}{n}\]is plugged in. This likelihood is now a function of only \(\beta\) and is technically referred to as a profile-likelihood.
References
[1]Green. “Econometric Analysis,” 5th ed., Pearson, 2003.
- predict(design=None)¶
After a model has been fit, results are (assumed to be) stored in self.results, which itself should have a predict method.
- rank()¶
Compute rank of design matrix
- score(beta, Y, nuisance=None)¶
Gradient of the loglikelihood function at (beta, Y, nuisance).
The graient of the loglikelihood function at (beta, Y, nuisance) is the score function.
See
logL()for details.- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- Yndarray
The dependent variable.
- nuisancedict, optional
A dict with key ‘sigma’, which is an optional estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / n, where n=Y.shape[0], X=self.design.
- Returns:
- The gradient of the loglikelihood function.
- whiten(Y)¶
Whiten design matrix
- Parameters:
- Xarray
design matrix
- Returns:
- wXarray
This matrix is the matrix whose pseudoinverse is ultimately used in estimating the coefficients. For OLSModel, it is does nothing. For WLSmodel, ARmodel, it pre-applies a square root of the covariance matrix to X.
OLSModel¶
- class nipy.algorithms.statistics.models.regression.OLSModel(design)¶
Bases:
LikelihoodModelA simple ordinary least squares model.
- Parameters:
- designarray-like
This is your design matrix. Data are assumed to be column ordered with observations in rows.
Examples
>>> from nipy.algorithms.statistics.api import Term, Formula >>> data = np.rec.fromarrays(([1,3,4,5,2,3,4], range(1,8)), ... names=('Y', 'X')) >>> f = Formula([Term("X"), 1]) >>> dmtx = f.design(data, return_float=True) >>> model = OLSModel(dmtx) >>> results = model.fit(data['Y']) >>> results.theta array([ 0.25 , 2.14285714]) >>> results.t() array([ 0.98019606, 1.87867287]) >>> print(results.Tcontrast([0,1])) <T contrast: effect=2.14285714286, sd=1.14062281591, t=1.87867287326, df_den=5> >>> print(results.Fcontrast(np.eye(2))) <F contrast: F=19.4607843137, df_den=5, df_num=2>
- Attributes:
- designndarray
This is the design, or X, matrix.
- wdesignndarray
This is the whitened design matrix. design == wdesign by default for the OLSModel, though models that inherit from the OLSModel will whiten the design.
- calc_betandarray
This is the Moore-Penrose pseudoinverse of the whitened design matrix.
- normalized_cov_betandarray
np.dot(calc_beta, calc_beta.T)- df_residscalar
Degrees of freedom of the residuals. Number of observations less the rank of the design.
- df_modelscalar
Degrees of freedome of the model. The rank of the design.
Methods
model.__init___(design)
model.logL(b=self.beta, Y)
- __init__(design)¶
- Parameters:
- designarray-like
This is your design matrix. Data are assumed to be column ordered with observations in rows.
- fit(Y)¶
Fit model to data Y
Full fit of the model including estimate of covariance matrix, (whitened) residuals and scale.
- Parameters:
- Yarray-like
The dependent variable for the Least Squares problem.
- Returns:
- fitRegressionResults
- has_intercept()¶
Check if column of 1s is in column space of design
- information(beta, nuisance=None)¶
Returns the information matrix at (beta, Y, nuisance).
See logL for details.
- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- nuisancedict
A dict with key ‘sigma’, which is an estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / nwhere n=Y.shape[0], X=self.design.
- Returns:
- infoarray
The information matrix, the negative of the inverse of the Hessian of the of the log-likelihood function evaluated at (theta, Y, nuisance).
- initialize(design)¶
Initialize (possibly re-initialize) a Model instance.
For instance, the design matrix of a linear model may change and some things must be recomputed.
- logL(beta, Y, nuisance=None)¶
Returns the value of the loglikelihood function at beta.
Given the whitened design matrix, the loglikelihood is evaluated at the parameter vector, beta, for the dependent variable, Y and the nuisance parameter, sigma.
- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- Yndarray
The dependent variable
- nuisancedict, optional
A dict with key ‘sigma’, which is an optional estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / n, where n=Y.shape[0], X=self.design.
- Returns:
- loglffloat
The value of the loglikelihood function.
Notes
The log-Likelihood Function is defined as
\[\ell(\beta,\sigma,Y)= -\frac{n}{2}\log(2\pi\sigma^2) - \|Y-X\beta\|^2/(2\sigma^2)\]The parameter \(\sigma\) above is what is sometimes referred to as a nuisance parameter. That is, the likelihood is considered as a function of \(\beta\), but to evaluate it, a value of \(\sigma\) is needed.
If \(\sigma\) is not provided, then its maximum likelihood estimate:
\[\hat{\sigma}(\beta) = \frac{\text{SSE}(\beta)}{n}\]is plugged in. This likelihood is now a function of only \(\beta\) and is technically referred to as a profile-likelihood.
References
[1]Green. “Econometric Analysis,” 5th ed., Pearson, 2003.
- predict(design=None)¶
After a model has been fit, results are (assumed to be) stored in self.results, which itself should have a predict method.
- rank()¶
Compute rank of design matrix
- score(beta, Y, nuisance=None)¶
Gradient of the loglikelihood function at (beta, Y, nuisance).
The graient of the loglikelihood function at (beta, Y, nuisance) is the score function.
See
logL()for details.- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- Yndarray
The dependent variable.
- nuisancedict, optional
A dict with key ‘sigma’, which is an optional estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / n, where n=Y.shape[0], X=self.design.
- Returns:
- The gradient of the loglikelihood function.
- whiten(X)¶
Whiten design matrix
- Parameters:
- Xarray
design matrix
- Returns:
- wXarray
This matrix is the matrix whose pseudoinverse is ultimately used in estimating the coefficients. For OLSModel, it is does nothing. For WLSmodel, ARmodel, it pre-applies a square root of the covariance matrix to X.
RegressionResults¶
- class nipy.algorithms.statistics.models.regression.RegressionResults(theta, Y, model, wY, wresid, cov=None, dispersion=1.0, nuisance=None)¶
Bases:
LikelihoodModelResultsThis class summarizes the fit of a linear regression model.
It handles the output of contrasts, estimates of covariance, etc.
- __init__(theta, Y, model, wY, wresid, cov=None, dispersion=1.0, nuisance=None)¶
See LikelihoodModelResults constructor.
The only difference is that the whitened Y and residual values are stored for a regression model.
- AIC()¶
Akaike Information Criterion
- BIC()¶
Schwarz’s Bayesian Information Criterion
- F_overall()¶
Overall goodness of fit F test, comparing model to a model with just an intercept. If not an OLS model this is a pseudo-F.
- Fcontrast(matrix, dispersion=None, invcov=None)¶
Compute an Fcontrast for a contrast matrix matrix.
Here, matrix M is assumed to be non-singular. More precisely
\[M pX pX' M'\]is assumed invertible. Here, \(pX\) is the generalized inverse of the design matrix of the model. There can be problems in non-OLS models where the rank of the covariance of the noise is not full.
See the contrast module to see how to specify contrasts. In particular, the matrices from these contrasts will always be non-singular in the sense above.
- Parameters:
- matrix1D array-like
contrast matrix
- dispersionNone or float, optional
If None, use
self.dispersion- invcovNone or array, optional
Known inverse of variance covariance matrix. If None, calculate this matrix.
- Returns:
- f_res
FContrastResultsinstance with attributes F, df_den, df_num
- f_res
Notes
For F contrasts, we now specify an effect and covariance
- MSE()¶
Mean square (error)
- MSR()¶
Mean square (regression)
- MST()¶
Mean square (total)
- R2()¶
Return the adjusted R^2 value for each row of the response Y.
Notes
Changed to the textbook definition of R^2.
See: Davidson and MacKinnon p 74
- R2_adj()¶
Return the R^2 value for each row of the response Y.
Notes
Changed to the textbook definition of R^2.
See: Davidson and MacKinnon p 74
- SSE()¶
Error sum of squares. If not from an OLS model this is “pseudo”-SSE.
- SSR()¶
Regression sum of squares
- SST()¶
Total sum of squares. If not from an OLS model this is “pseudo”-SST.
- Tcontrast(matrix, store=('t', 'effect', 'sd'), dispersion=None)¶
Compute a Tcontrast for a row vector matrix
To get the t-statistic for a single column, use the ‘t’ method.
- Parameters:
- matrix1D array-like
contrast matrix
- storesequence, optional
components of t to store in results output object. Defaults to all components (‘t’, ‘effect’, ‘sd’).
- dispersionNone or float, optional
- Returns:
- res
TContrastResultsobject
- res
- conf_int(alpha=0.05, cols=None, dispersion=None)¶
The confidence interval of the specified theta estimates.
- Parameters:
- alphafloat, optional
The alpha level for the confidence interval. ie., alpha = .05 returns a 95% confidence interval.
- colstuple, optional
cols specifies which confidence intervals to return
- dispersionNone or scalar
scale factor for the variance / covariance (see class docstring and
vcovmethod docstring)
- Returns:
- cisndarray
cis is shape
(len(cols), 2)where each row contains [lower, upper] for the given entry in cols
Notes
Confidence intervals are two-tailed. TODO: tails : string, optional
tails can be “two”, “upper”, or “lower”
Examples
>>> from numpy.random import standard_normal as stan >>> from nipy.algorithms.statistics.models.regression import OLSModel >>> x = np.hstack((stan((30,1)),stan((30,1)),stan((30,1)))) >>> beta=np.array([3.25, 1.5, 7.0]) >>> y = np.dot(x,beta) + stan((30)) >>> model = OLSModel(x).fit(y) >>> confidence_intervals = model.conf_int(cols=(1,2))
- logL()¶
The maximized log-likelihood
- norm_resid()¶
Residuals, normalized to have unit length.
Notes
Is this supposed to return “standardized residuals,” residuals standardized to have mean zero and approximately unit variance?
d_i = e_i / sqrt(MS_E)
Where MS_E = SSE / (n - k)
- See: Montgomery and Peck 3.2.1 p. 68
Davidson and MacKinnon 15.2 p 662
- predicted()¶
Return linear predictor values from a design matrix.
- resid()¶
Residuals from the fit.
- t(column=None)¶
Return the (Wald) t-statistic for a given parameter estimate.
Use Tcontrast for more complicated (Wald) t-statistics.
- vcov(matrix=None, column=None, dispersion=None, other=None)¶
Variance/covariance matrix of linear contrast
- Parameters:
- matrix: (dim, self.theta.shape[0]) array, optional
numerical contrast specification, where
dimrefers to the ‘dimension’ of the contrast i.e. 1 for t contrasts, 1 or more for F contrasts.- column: int, optional
alternative way of specifying contrasts (column index)
- dispersion: float or (n_voxels,) array, optional
value(s) for the dispersion parameters
- other: (dim, self.theta.shape[0]) array, optional
alternative contrast specification (?)
- Returns:
- cov: (dim, dim) or (n_voxels, dim, dim) array
the estimated covariance matrix/matrices
- Returns the variance/covariance matrix of a linear contrast of the
- estimates of theta, multiplied by dispersion which will often be an
- estimate of dispersion, like, sigma^2.
- The covariance of interest is either specified as a (set of) column(s)
- or a matrix.
WLSModel¶
- class nipy.algorithms.statistics.models.regression.WLSModel(design, weights=1)¶
Bases:
OLSModelA regression model with diagonal but non-identity covariance structure.
The weights are presumed to be (proportional to the) inverse of the variance of the observations.
Examples
>>> from nipy.algorithms.statistics.api import Term, Formula >>> data = np.rec.fromarrays(([1,3,4,5,2,3,4], range(1,8)), ... names=('Y', 'X')) >>> f = Formula([Term("X"), 1]) >>> dmtx = f.design(data, return_float=True) >>> model = WLSModel(dmtx, weights=range(1,8)) >>> results = model.fit(data['Y']) >>> results.theta array([ 0.0952381 , 2.91666667]) >>> results.t() array([ 0.35684428, 2.0652652 ]) >>> print(results.Tcontrast([0,1])) <T contrast: effect=2.91666666667, sd=1.41224801095, t=2.06526519708, df_den=5> >>> print(results.Fcontrast(np.identity(2))) <F contrast: F=26.9986072423, df_den=5, df_num=2>
- __init__(design, weights=1)¶
- Parameters:
- designarray-like
This is your design matrix. Data are assumed to be column ordered with observations in rows.
- fit(Y)¶
Fit model to data Y
Full fit of the model including estimate of covariance matrix, (whitened) residuals and scale.
- Parameters:
- Yarray-like
The dependent variable for the Least Squares problem.
- Returns:
- fitRegressionResults
- has_intercept()¶
Check if column of 1s is in column space of design
- information(beta, nuisance=None)¶
Returns the information matrix at (beta, Y, nuisance).
See logL for details.
- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- nuisancedict
A dict with key ‘sigma’, which is an estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / nwhere n=Y.shape[0], X=self.design.
- Returns:
- infoarray
The information matrix, the negative of the inverse of the Hessian of the of the log-likelihood function evaluated at (theta, Y, nuisance).
- initialize(design)¶
Initialize (possibly re-initialize) a Model instance.
For instance, the design matrix of a linear model may change and some things must be recomputed.
- logL(beta, Y, nuisance=None)¶
Returns the value of the loglikelihood function at beta.
Given the whitened design matrix, the loglikelihood is evaluated at the parameter vector, beta, for the dependent variable, Y and the nuisance parameter, sigma.
- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- Yndarray
The dependent variable
- nuisancedict, optional
A dict with key ‘sigma’, which is an optional estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / n, where n=Y.shape[0], X=self.design.
- Returns:
- loglffloat
The value of the loglikelihood function.
Notes
The log-Likelihood Function is defined as
\[\ell(\beta,\sigma,Y)= -\frac{n}{2}\log(2\pi\sigma^2) - \|Y-X\beta\|^2/(2\sigma^2)\]The parameter \(\sigma\) above is what is sometimes referred to as a nuisance parameter. That is, the likelihood is considered as a function of \(\beta\), but to evaluate it, a value of \(\sigma\) is needed.
If \(\sigma\) is not provided, then its maximum likelihood estimate:
\[\hat{\sigma}(\beta) = \frac{\text{SSE}(\beta)}{n}\]is plugged in. This likelihood is now a function of only \(\beta\) and is technically referred to as a profile-likelihood.
References
[1]Green. “Econometric Analysis,” 5th ed., Pearson, 2003.
- predict(design=None)¶
After a model has been fit, results are (assumed to be) stored in self.results, which itself should have a predict method.
- rank()¶
Compute rank of design matrix
- score(beta, Y, nuisance=None)¶
Gradient of the loglikelihood function at (beta, Y, nuisance).
The graient of the loglikelihood function at (beta, Y, nuisance) is the score function.
See
logL()for details.- Parameters:
- betandarray
The parameter estimates. Must be of length df_model.
- Yndarray
The dependent variable.
- nuisancedict, optional
A dict with key ‘sigma’, which is an optional estimate of sigma. If None, defaults to its maximum likelihood estimate (with beta fixed) as
sum((Y - X*beta)**2) / n, where n=Y.shape[0], X=self.design.
- Returns:
- The gradient of the loglikelihood function.
- whiten(X)¶
Whitener for WLS model, multiplies by sqrt(self.weights)
Functions¶
- nipy.algorithms.statistics.models.regression.ar_bias_correct(results, order, invM=None)¶
Apply bias correction in calculating AR(p) coefficients from results
There is a slight bias in the rho estimates on residuals due to the correlations induced in the residuals by fitting a linear model. See [Worsley2002].
This routine implements the bias correction described in appendix A.1 of [Worsley2002].
- Parameters:
- resultsndarray or results object
If ndarray, assume these are residuals, from a simple model. If a results object, with attribute
resid, then use these for the residuals. See Notes for more detail- orderint
Order
pof AR(p) model- invMNone or array
Known bias correcting matrix for covariance. If None, calculate from
results.model
- Returns:
- rhoarray
Bias-corrected AR(p) coefficients
Notes
If results has attributes
residandscale, then assumescalehas come from a fit of a potentially customized model, and we use that for the sum of squared residuals. In this case we also needresults.df_resid. Otherwise we assume this is a simple Gaussian model, like OLS, and take the simple sum of squares of the residuals.References
- nipy.algorithms.statistics.models.regression.ar_bias_corrector(design, calc_beta, order=1)¶
Return bias correcting matrix for design and AR order order
There is a slight bias in the rho estimates on residuals due to the correlations induced in the residuals by fitting a linear model. See [Worsley2002].
This routine implements the bias correction described in appendix A.1 of [Worsley2002].
- Parameters:
- designarray
Design matrix
- calc_betaarray
Moore-Penrose pseudoinverse of the (maybe) whitened design matrix. This is the matrix that, when applied to the (maybe whitened) data, produces the betas.
- orderint, optional
Order p of AR(p) process
- Returns:
- invMarray
Matrix to bias correct estimated covariance matrix in calculating the AR coefficients
References
- nipy.algorithms.statistics.models.regression.isestimable(C, D)¶
True if (Q, P) contrast C is estimable for (N, P) design D
From an Q x P contrast matrix C and an N x P design matrix D, checks if the contrast C is estimable by looking at the rank of
vstack([C,D])and verifying it is the same as the rank of D.- Parameters:
- C(Q, P) array-like
contrast matrix. If C has is 1 dimensional assume shape (1, P)
- D: (N, P) array-like
design matrix
- Returns:
- tfbool
True if the contrast C is estimable on design D
Examples
>>> D = np.array([[1, 1, 1, 0, 0, 0], ... [0, 0, 0, 1, 1, 1], ... [1, 1, 1, 1, 1, 1]]).T >>> isestimable([1, 0, 0], D) False >>> isestimable([1, -1, 0], D) True
- nipy.algorithms.statistics.models.regression.yule_walker(X, order=1, method='unbiased', df=None, inv=False)¶
Estimate AR(p) parameters from a sequence X using Yule-Walker equation.
unbiased or maximum-likelihood estimator (mle)
See, for example:
http://en.wikipedia.org/wiki/Autoregressive_moving_average_model
- Parameters:
- Xndarray of shape(n)
- orderint, optional
Order of AR process.
- methodstr, optional
Method can be “unbiased” or “mle” and this determines denominator in estimate of autocorrelation function (ACF) at lag k. If “mle”, the denominator is n=X.shape[0], if “unbiased” the denominator is n-k.
- dfint, optional
Specifies the degrees of freedom. If df is supplied, then it is assumed the X has df degrees of freedom rather than n.
- invbool, optional
Whether to return the inverse of the R matrix (see code)
- Returns:
- rho(order,) ndarray
- sigmaint
standard deviation of the residuals after fit
- R_invndarray
If inv is True, also return the inverse of the R matrix
Notes
See also http://en.wikipedia.org/wiki/AR_model#Calculation_of_the_AR_parameters
